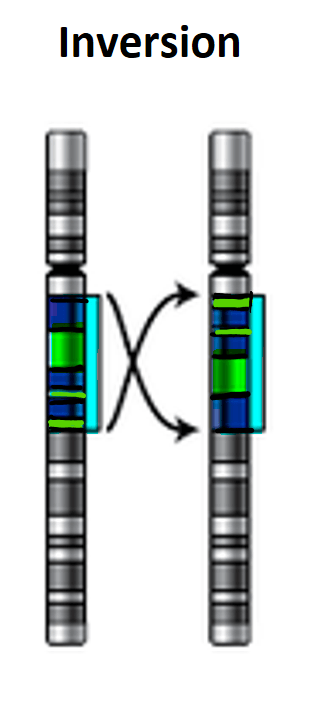
What are chromosome inversions?
Inversions are a diverse class of chromosomal mutation. Most are small (<1KB). Others, for example, the famous 3RP inversion of Drosophila melanogaster, are several megabases in size, including several percentages of the entire genome, and encompass hundreds or thousands of genes. Inversions fall into two classes: pericentric inversions include a centromere, while paracentric inversions do not. With pericentric inversions, a single crossover event that occurs between the breakpoints of a heterozygote produces unbalanced gametes that carry deletions, insertions, and zero or two centromeres.
This can reduce fertility, making the inversions subdominant (decreased heterozygous fitness). However, some pericentric inversions apparently escape fitness costs when heterozygous, perhaps because they somehow suppress recombination. Although these may represent only a small fraction of all pericentric inversions that arise by mutation, they are likely to be highly enriched among those that spread to fixation. There are large systematic differences between taxa in the frequency and severity of fitness effects.
For example, heterozygotes for inversions appear to show decreased fertility in plants much more frequently than in animals. In contrast, many of the paracentric inversions that segregate in nature may not suffer from under dominance. This is probably one of the main reasons why they are orders of magnitude more common than pericentric inversions, both as internal polymorphisms and as fixed differences between species. Inversions were first observed in the giant salivary chromosomes of fly larvae, and Diptera remains the group in which large inversions can most easily be detected.
Chromosome staining techniques can visualize inversions in some other groups, including mammals, but with much lower resolution (and higher effort). The presence of an inversion is suggested when a certain cross consistently shows blocked recombination in part of the genome, but this observation requires genetic markers that have been mapped. Sequencing is a third way in which inversions are detected. The short reads that are characteristic of current high-throughput sequencing methods are adequate for determining whether an individual has an inversion that has already been characterized by its breakpoints, but this technology is poor at searching for new inversions.
Inversions and Recombination
A key evolutionary effect of inversions is that they suppress recombination as heterozygotes. Suppression stems from the loss of unbalanced gametes resulting from recombination, failure of inverted regions to synapse in heterozygotes, and probably other mechanisms not yet understood. Large inversions show very low (but still positive) recombination rates as heterozygotes, resulting from double crossovers and gene conversion, but the rates are orders of magnitude smaller than homozygotes. However, on timescales of 105 generations and beyond, even this very limited recombination complements mutation as a major source of genetic variation within inversions, which originate from a single chromosome and thus have no variation. some when they first appear.
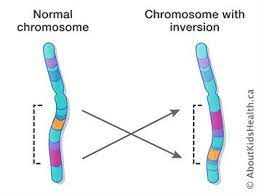
One way to visualize the evolutionary properties of investments is through an analogy. Consider the parallelism between populations of inverted and non-inverted chromosomes, on the one hand, and a pair of coexisting biological species on the other. Within each of the two species, the normal rules of Mendelian inheritance apply. Between them, however, there is little genetic exchange, except for an occasional hybridization event (such as the rare recombination between inverted and non-inverted chromosomes). Ecological competition between the two species (such as fitness differences between the two chromosomal forms) may result in the coexistence of the two species (as a stable polymorphism) or in the replacement of one species by another (such as fixation of the ancestral chromosomal or inverted) form).
How do inversions evolve?
Like other types of mutations, inversions evolve under selection and random drift. Many inversions, particularly small ones in intergenic regions, are likely to evolve neutrally (drift only). Selection can result in three ways. Inversions can cause structural problems with meiosis, as is the case with some pericentric inversions. Alternatively, a breakpoint can interrupt an open reading frame or alter gene expression. The consequences can be detrimental, as in some human genetic diseases, but in other cases, it could lead to adaptive mutation. Finally, selection can act on an inversion when it carries one or more selected alleles.
Many pericentric inversions are subdominant, posing an evolutionary puzzle. It is selected against a subdominant inversion, as long as it is rare. However, closely related species often show fixed differences, implying that an inversion must nevertheless have appeared and spread through one of the two lineages from their last common ancestor. Some researchers have invoked drift to solve this puzzle. A line of support for this hypothesis is the observation that annual plants show high rates of evolution of subdominant chromosomal rearrangements. Many annual plants have large population fluctuations and, at least occasionally, self-fertilize, dramatically reducing effective population size and thus increasing the power of drift.
Conversely, inversions can also be overdominant (superior as heterozygotes). The genetic basis of the overdominance appears not to have been determined for any inversion. In principle, it could result from the effects of breakpoints. Alternatively, it could be the result of overdominance at a locus within the inversion if one allele fixates on the ancestral chromosome and the other allele fixes on the inverted chromosome. A third hypothesis is “associative overdominance”. This occurs when an inversion occurs to capture one or more deleterious recessive alleles, which is likely to make a large inversion.
If inversion is selectively favoured when rare, it may extend to the point where homozygous recessives become frequent enough to offset the initial advantage. The result is a balanced polymorphism that has the same evolutionary properties as conventional overdominance. Some inversions show meiotic drive: gametes from heterozygotes carry the inversion more than 50% of the time. Meiotic drive systems often involve a pair of interacting loci that must be inherited for the system to invade a population.
An inversion can suppress recombination that would otherwise disrupt the drive system, and then spreads as the drive alleles spread. One reason evolutionary biologists are fascinated with inversions is that they are highly polymorphic in some species. Polymorphic inversions do not appear to be ancient, at least in flies, with ages on the order of 106 generations. An intriguing mystery is why there are huge differences in polymorphism levels and inversion fixation rates between closely related species and between chromosomes within a species, for example in Drosophila.